-Search query
-Search result
Showing 1 - 50 of 203 items for (author: hansen & g)

EMDB-41363: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5
Method: single particle / : Yue H, Hunkeler M, Roy Burman SS, Fischer ES

PDB-8tl6: 
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5
Method: single particle / : Yue H, Hunkeler M, Roy Burman SS, Fischer ES

EMDB-18207: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide, Glacios map
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-18230: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA
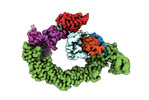
EMDB-18915: 
Ubiquitin ligation to neosubstrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB-BRD4 BD2
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

PDB-8q7r: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

PDB-8r5h: 
Ubiquitin ligation to neosubstrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB-BRD4 BD2
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-17412: 
Vaccinia Virus flower-shaped pore-like structure
Method: single particle / : Hansen J, Datler J, Thader A, Schloegl A, Hodirnau VV, Schur FKM

EMDB-40984: 
5TU-t1 - heterodimeric triplet polymerase ribozyme
Method: single particle / : McRae EKS, Kristoffersen E, Gallego I, Hansen K, Holliger P, Andersen ES

PDB-8t2p: 
5TU-t1 - heterodimeric triplet polymerase ribozyme
Method: single particle / : McRae EKS, Kristoffersen E, Gallego I, Hansen K, Holliger P, Andersen ES

EMDB-17410: 
Vaccinia Virus palisade layer A10 trimer
Method: single particle / : Datler J, Hansen JM, Thader A, Schloegl A, Hodirnau VV, Schur FKM

EMDB-17411: 
Subtomogram averaging structure of the Vaccinia Virus core palisade layer
Method: subtomogram averaging / : Datler J, Hansen JM, Thader A, Schloegl A, Hodirnau VV, Schur FKM

EMDB-17413: 
Cryo-electron tomogram of intact mature Vaccinia Virus particle
Method: electron tomography / : Datler J, Hansen JM, Thader A, Schloegl A, Hodirnau VV, Schur FKM
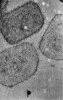
EMDB-17414: 
Cryo-electron tomogram of isolated Vaccinia Virus cores
Method: electron tomography / : Datler J, Hansen JM, Thader A, Schloegl A, Hodirnau VV, Schur FKM

EMDB-18452: 
Vaccinia virus inner core wall
Method: single particle / : Datler J, Hansen JM, Thader A, Schloegl A, Hodirnau VV, Schur FKM

PDB-8p4k: 
Vaccinia Virus palisade layer A10 trimer
Method: single particle / : Datler J, Hansen JM, Thader A, Schloegl A, Hodirnau VV, Schur FKM

EMDB-16024: 
SARS-CoV-2 S protein in complex with pT1644 Fab
Method: single particle / : Stroeh L, Hansen G, Vollmer B, Krey T, Benecke T, Gruenewald K

EMDB-16026: 
SARS-CoV-2 S protein in complex with pT1696 Fab
Method: single particle / : Hansen G, Benecke T, Vollmer B, Gruenewald K, Krey T

PDB-8bg6: 
SARS-CoV-2 S protein in complex with pT1644 Fab
Method: single particle / : Stroeh L, Hansen G, Vollmer B, Krey T, Benecke T, Gruenewald K

PDB-8bg8: 
SARS-CoV-2 S protein in complex with pT1696 Fab
Method: single particle / : Hansen G, Benecke T, Vollmer B, Gruenewald K, Krey T

EMDB-16416: 
HB3VAR03 apo headstructure (PfEMP1 A) complexed with EPCR
Method: single particle / : Raghavan SSR, Lavstsen T, Wang KT

PDB-8c44: 
HB3VAR03 apo headstructure (PfEMP1 A) complexed with EPCR
Method: single particle / : Raghavan SSR, Lavstsen T, Wang KT

EMDB-27920: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

EMDB-27921: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

PDB-8e6j: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

PDB-8e6k: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

EMDB-16415: 
HB3VAR03 apo headstructure (PfEMP1 A)
Method: single particle / : Raghavan SSR, Lavstsen T, Wang KT

PDB-8c3y: 
HB3VAR03 apo headstructure (PfEMP1 A)
Method: single particle / : Raghavan SSR, Lavstsen T, Wang KT

EMDB-27738: 
Negative stain EM map of the heterodimeric p110gamma-p84 complex
Method: single particle / : Burke JE, Dalwadi U, Rathinaswamy MK, Yip CK, Nam SE

EMDB-27627: 
Structure of p110 gamma bound to the Ras inhibitory nanobody NB7
Method: single particle / : Burke JE, Nam SE, Rathinaswamy MK, Yip CK

PDB-8dp0: 
Structure of p110 gamma bound to the Ras inhibitory nanobody NB7
Method: single particle / : Burke JE, Nam SE, Rathinaswamy MK, Yip CK

EMDB-29019: 
Alpha1/BetaB Heteromeric Glycine Receptor in 1 mM Glycine 20 uM Ivermectin State
Method: single particle / : Gibbs E, Chakrapani S

PDB-8fe1: 
Alpha1/BetaB Heteromeric Glycine Receptor in 1 mM Glycine 20 uM Ivermectin State
Method: single particle / : Gibbs E, Chakrapani S

EMDB-26989: 
Extracellular architecture of an engineered canonical Wnt signaling ternary complex
Method: single particle / : Tsutsumi N, Jude KM, Garcia KC

PDB-8ctg: 
Extracellular architecture of an engineered canonical Wnt signaling ternary complex
Method: single particle / : Tsutsumi N, Jude KM, Garcia KC

EMDB-25699: 
VFLIP Spike Trimer with GAR03
Method: single particle / : Sobti M, Stewart AG

EMDB-25700: 
VFLIP Spike Trimer with GAR05 FAB
Method: single particle / : Sobti M, Stewart AG, Rouet R, Langley DB

PDB-7t5o: 
VFLIP Spike Trimer with GAR03
Method: single particle / : Sobti M, Stewart AG, Rouet R, Langley DB

EMDB-25575: 
D8-C4 computationally-designed Rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25576: 
D3-C5 computationally-designed rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D
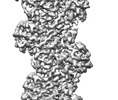
EMDB-26987: 
T148-ADPR-F-actin
Method: helical / : Egelman EH

EMDB-25824: 
aRML prion fibril
Method: single particle / : Hoyt F, Standke HG, Artikis E, Caughey B, Kraus A
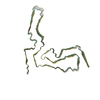
PDB-7td6: 
aRML prion fibril
Method: single particle / : Hoyt F, Standke HG, Artikis E, Caughey B, Kraus A

EMDB-13326: 
T. maritima CorA in DDM micelles without Mg2+ bound in D2O
Method: single particle / : Larsen AH, Johansen NT, Arleth L

EMDB-13327: 
T. maritima CorA in DDM micelles with Mg2+ bound in D2O
Method: single particle / : Larsen AH, Johansen NT, Arleth L

EMDB-23812: 
Structure of the apo phosphoinositide 3-kinase p110 gamma (PIK3CG) p101 (PIK3R5) complex
Method: single particle / : Burke JE, Dalwadi U, Rathinaswamy MK, Yip CK

EMDB-25578: 
D3-C5 computationally-designed rotor (axel only)
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25579: 
D3-C3 computationally-designed rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-25580: 
C3-C3 computationally-designed rotor
Method: single particle / : Hansen JM, Courbet A, Quispe J, Kollman JM, Baker D

EMDB-23831: 
Human CTPS1 bound to UTP, AMPPNP, and glutamine
Method: single particle / : Lynch EM, Dimattia MA, Kollman JM
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
